Published on 21.04.2023
Six Main Tasks in Image Processing

Applied Imaging Processing – Concepts and Techniques
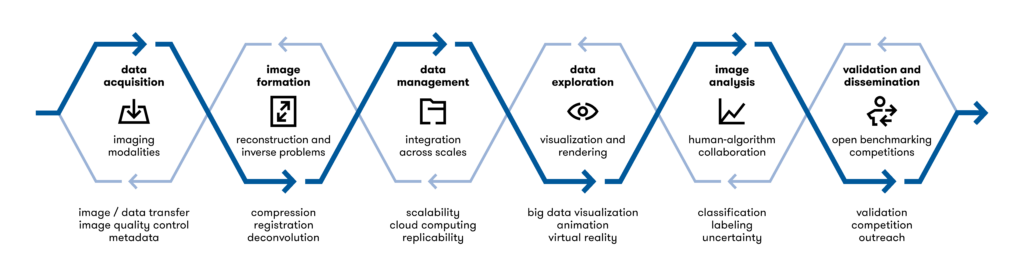
In this series of (virtual) seminars, six key image processing tasks will be discussed, following a typical workflow in the image processing pipeline.
Images are not always captured by a camera. Often, they must be tediously reconstructed from a series of projections or other non-image types of acquisitions. Different reconstruction algorithms allow for better image quality or can focus on specific properties of the objects under observation. Noise can be introduced at many steps in the image acquisition process. Denoising is therefore an essential step in most image processing workflows. Tracking individual objects over multiple time steps is a difficult task, but allows for the observation of temporal dynamics. Segmentation refers to the assignment of each pixel in an image to a specific category. In semantic segmentation, all pixels belonging to a cat are labeled “cat”, and all pixels belonging to trees are labeled “tree”. In instance segmentation, each pixel is additionally assigned to an object instance, making it possible to distinguish multiple cats and trees in an image. The visualization of otherwise difficult-to-interpret data, such as reconstructed 3D(+T) objects or high-dimensional image data, is essential for understanding the results. Finally, interpreting the results of AI-based image analysis algorithms is important: Why was a particular decision made? What structures in the images were responsible? What can AI tell us about the underlying problem?
This seminar series is held by imaging experts from and invited by the organizing schools and Helmholtz Imaging. The course consists of lectures and interactive discussions. It covers various image processing techniques used in life sciences and soft matter. It is recommended to attend all lectures for a deep understanding of the subject. Registration is required to attend. Places are limited and priority will be given to fellows (members) of the three schools in case of overbooking. Participants with an attendance rate of more than 70% may receive a certificate of attendance.
Join this seminar series! More information: events.hifis.net/event/793
Registration is required; a registration link can be requested via mail to sabine.niebel@helmholtz-imaging.de.
Overview of seminars
4 May 2023: Introduction
The six main tasks in image processing: an overview
Modern imaging methods enable us to capture structure and dynamics in unprecedented detail and with high temporal and spatial resolution. The challenge is to make full use of the potential of such “big” imaging data to obtain quantitative results, test hypotheses, and develop new theories and models. Manual or semi-automated image analysis workflows quickly become a bottleneck because they do not scale well with the amount and complexity of the data. This lecture provides an overview of typical automated image analysis workflows using examples from high-resolution microscopy in the life sciences. We will cover the six main tasks, including image reconstruction from raw tomography or localization data, denoising, tracking in time and space, segmentation to extract objects from images, visualization, and explainable AI-based methods. In the last part, we will introduce tools for integrating these different tasks into complete workflows.
Speaker: Prof. Philip Kollmannsberger (Heinrich-Heine-Universität Düsseldorf)
25 May 2023: Reconstruction & Denoising
Tomographic methods in medical imaging
In medical imaging, dedicated cameras are used to capture images, and medical images are typically three-dimensional images. These three-dimensional images are generated from a series of datagrams or images of sections and/or projections, which is achieved by tomographic reconstruction. Magnetic Resonance Imaging (MRI), Computed Tomography (CT), Positron Emission Tomography (PET) and Single Photon Emission Computed Tomography (SPECT) are typical examples of different imaging modalities where tomographic reconstruction methods are used. This lecture gives a brief introduction to Fourier transform based reconstruction (MRI), filtered back projection (CT) and iterative reconstruction (PET and CT).
Speaker: Dr. Christoph Lerche (PET Physics Group at Forschungszentrum Jülich)
Restoring noisy microscopy images
Bleaching and phototoxicity in light microscopy, or beam-induced sample damage in electron microscopy, are just some of the many reasons why exposure must be kept low in microscopy. However, limiting exposure can result in noisy image acquisition. These noisy image data are not only tedious to browse, but often make automated image processing difficult or even impossible. In this talk, I will give an overview of different deep learning based image restoration methods that enable high quality image restoration for very noisy image data.
Speaker: Dr. Tim-Oliver Buchholz (Friedrich Miescher Institute for Biomedical Research, Basel)
15 June 2023: Tracking
Tracking of objects: from one to many
I will talk about two separate research areas for tracking objects over time and instances. The first applies to the scenario of tracking an object over time, e.g. a known 3D rigid model. I will introduce Bayesian and particle filters and explain some technical ideas to make them fast and accurate. In the second part of the lecture I will introduce the field of tracking a large set of objects (e.g. cells) over time, and also instances. Since the objects typically do not move randomly, it makes sense to formalize their “structured motion” and formulate this task as a structured prediction problem. To this end, I will introduce efficient solvers for this problem.
Speaker: Prof. Carsten Rother (HCI, Universität Heidelberg)
29 June 2023: Segmentation
Machine Learning for Image Segmentation
The course will introduce current machine learning models for image segmentation, including semantic- and instance segmentation. It will also cover how to apply such models to large data. A respective tutorial at the end of the seminar series will give hands-on experience in training and applying such models.
Speaker: Prof. Dagmar Kainmüller (Helmholtz Imaging, Max-Delbrück Center, Berlin)
6 July 2023: Visualization
Visualizing spatial datasets
Join us in our journey through selected methods of telling the story of your dataset through visualizations. In the seminar, we will transform 3D segmentations into effectful Blender renderings.
Speaker: Deborah Schmidt (Helmholtz Imaging, Max-Delbrück Center, Berlin)
27 July 2023: Explainable Machine Learning
Explainable Machine Learning
Explainable machine learning involves two complementary types of questions: (1) Method designers and theorists ask: “Why do deep neural networks work so well? How can we further improve them? How can we provide formal performance guarantees?” (2) Instead, method users ask, “How did the network arrive at its conclusion? What variables determine the result, and in what way? Can the result be trusted?” The talk will introduce both areas and review the current state of our answers.
Speaker: Prof. Ullrich Köthe (Interdisciplinary Center for Scientific Computing (IWR) University of Heidelberg)
This event is organized by the International Helmholtz Research School of Biophysics and Soft Matter, BioInterfaces International Graduate School, Helmholtz Information & Data Science School for Health and Helmholtz Imaging.
